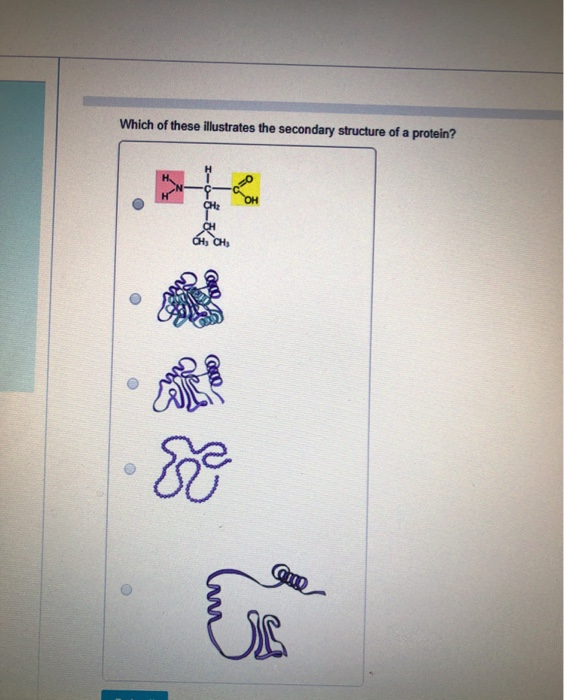
 
A really long helix can then produce a significant electric field and affect protein binding properties. This leads effectively to one long dipole of magnitude n(3.5) Debyes, where n is the number of amino acids in the helix. In the alpha helix al the amides in the backbone point in one direction from the C-terminus to N-terminus. It is the first instance of universal quantum algorithms beingĮmployed to tackle RNA folding problems, and our work provides a valuable modelįor utilizing universal quantum computers in solving RNA folding problems.\): Helix Dipole ( reproduce image) Sequences, our quantum approach outperforms classical energy-based methods.Įxperimental results also indicate our method is robust in current noisyĭevices. Throughout our benchmark dataset, simulation results suggest that our quantumĪpproach is comparable in accuracy to classical methods. For brevity, the 400 components in eqn 2 are denoted by y 1, y 2,, y 400. Performance with both numerical simulation and experimental realization. Solved through quantum approximation optimization algorithm. The state of a quadratic Hamiltonian, the whole folding process is described asĪ quadratic unconstrained binary optimization model. This resource integrates structural information from the RCSB PDB and protein positional features (such as sites of mutation, catalytic residues, and secondary. Mapped all possible secondary structure to The 3D Protein Feature View (PFV) maps protein sequence features ( i.e., 1D protein positions of interest) onto the 3D structure of protein assemblies for visualization and analysis. :max_bytes(150000):strip_icc()/protein_structures-580535ab3df78cbc28dbaec4.jpg)
Quantum algorithms will be presented, which are highly flexible and can beĪpplied to various physical devices. Universal quantum computing are not available. 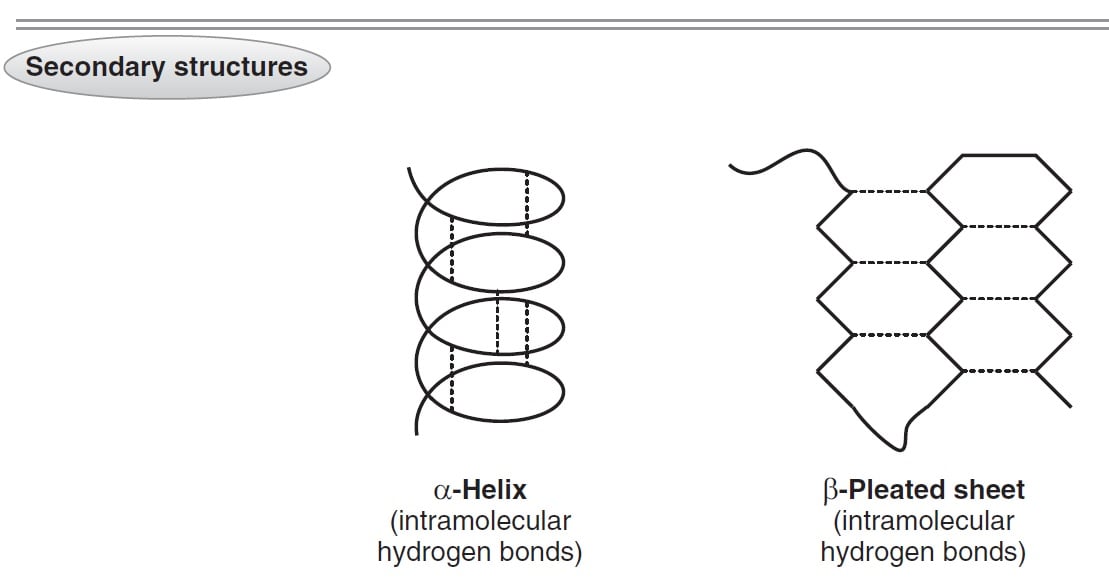
The folding of the secondary structure, highlighting that quantum computing isĪ promising solution to this problem. Recently, quantum annealer has rapidly predicted Sequences, while learning-based algorithms face challenges in obtaining TraditionalĮnergy-based algorithms are short of precision, particularly for non-nested Sequences that to know how its secondary structure is formed. Download a PDF of the paper titled Predicting RNA Secondary Structure on Universal Quantum Computer, by Ji Jiang and 7 other authors Download PDF Abstract: It is the first step for understanding how RNA structure folds from base 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
